Abstract:
Reconstructing the evolutionary tree for a set of 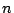 species
based on pairwise distances between the species is a fundamental problem in
bioinformatics. Neighbor joining is a popular distance based tree
reconstruction method. It always proposes fully resolved binary trees despite
missing evidence in the underlying distance data. Distance based methods
based on the theory of Buneman trees and refined Buneman trees avoid this
problem by only proposing evolutionary trees whose edges satisfy a number of
constraints. These trees might not be fully resolved but there is strong
combinatorial evidence for each proposed edge. The currently best algorithm
for computing the refined Buneman tree from a given distance measure has a
running time of
species
based on pairwise distances between the species is a fundamental problem in
bioinformatics. Neighbor joining is a popular distance based tree
reconstruction method. It always proposes fully resolved binary trees despite
missing evidence in the underlying distance data. Distance based methods
based on the theory of Buneman trees and refined Buneman trees avoid this
problem by only proposing evolutionary trees whose edges satisfy a number of
constraints. These trees might not be fully resolved but there is strong
combinatorial evidence for each proposed edge. The currently best algorithm
for computing the refined Buneman tree from a given distance measure has a
running time of 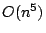 and a space consumption of
and a space consumption of 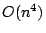 . In this paper,
we present an algorithm with running time
. In this paper,
we present an algorithm with running time 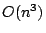 and space
consumption
and space
consumption 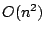
Available as PostScript, PDF, DVI.